DNA tests from the big consumer DNA companies like Ancestry.com and 23andMe rely on bead array technology.
This technology was invented in the 1990s by David Walt at Tufts University and further developed by Illumina, now a giant in the biotech world.
This article explains bead array technology in simple terms. A vague memory of high school science is all you need. Before we get to the bead arrays, we’ll start with some basic questions
What Is An Oligo?
“Oligo” is a friendly term for “oligonucleotide”. It’s certainly easier to pronounce.
An oligo is a short piece of DNA (or RNA) that is made in a laboratory.
When we manufacture DNA, we use the four building blocks of DNA known as A, C, T, and G.
Oligos are made by starting with one of these four DNA bases. Another base is added to it, and then another, and so on to form a chain in a known sequence.
When the oligo is complete, it has about twenty to thirty-five bases.

What Are Oligos Used For?
Oligos are used for lots of different purposes.
One example is a technique known as PCR or polymerase chain reaction. PCR turns one strand of DNA into millions of copies.
This means that analysis of a target fragment of DNA can be done by many machines in parallel. This revolutionized DNA analysis by greatly reducing the time and cost of DNA sequencing.
We’ll cover PCR elsewhere.
The rest of this article explains another use: to make oligonucleotide probes.
What Is An Oligonucleotide Probe?
You are probably familiar with the double-stranded helix structure of DNA.
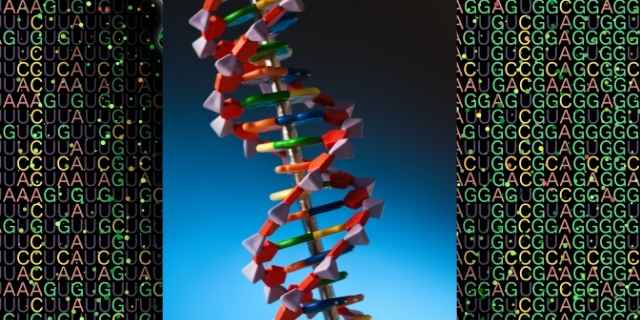
The rungs of the ladder are made by the building blocks of DNA on each strand that pair with each other.
These base units are nucleotides that we label A, C, T, and G:
- Adenine
- Cytosine
- Thymine
- Guanine
Suppose the sequence in one section of a single strand is “A T T C G A”.
Now we know what the sequence must be at the same positions in the other strand! That’s because DNA nucleotides only pair with one other type.
A only pairs with T and C only pairs with G.
So, we know that the complementary strand must be “T A A G C T”.
This knowledge underpins how probes work.
How do oligo probes work?
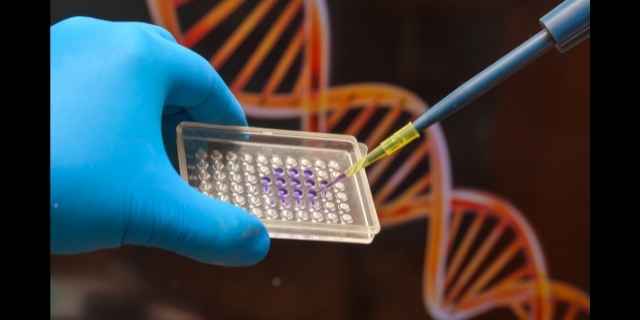
Let’s say that we’ve isolated a single strand of DNA in a laboratory dish and we want to figure out the nucleotide sequence.
We make a single-stranded oligo with the sequence “T A A G C T” and drop it into a dish with our sample. Then we add a few bits of chemical magic to make our oligo find its complementary sequence in the sample DNA.
The oligo has been treated with a fluorescent dye that makes each of the four base units light up in a different color.
When the oligo probe pairs up with its complementary sequence, the colors show us exactly where the “A T T C G A” sequence occurs in the sample.
What Are Bead Arrays?
There are different types of bead arrays so I’ll describe the version developed first by David Walt and then by the company he co-founded.
If you want to read more about the early days of a biotech giant, we have an article on Walt and the other co-founders of Illumina.
The technique uses optical fiber in an ingenious way. A tiny well is poked into the tip of 50 thousand optical fibers. The fibers are bundled together to form an array.
A tiny silica bead is placed into each well. The beads are almost “poured” across the fiber bundle, so they drop into their wells at random.
Every bead has an attached oligo probe that identifies a specific sequence of DNA.
So, now we have an array of beads with oligo probes.
The next step is to bring the target DNA sample to the array and let the probes bind to their complementary sequences in the sample.
Decoding the random array
At this point, the scientists don’t know which bead is in which well. As I mentioned, adding the beads to the wells is a random process.
The final step is to decode each bead i.e. figure out which oligo sequence is attached to which bead.
This is where things get a little confusing, but we won’t dwell on it.
Another set of oligo “decoder” probes is used to work out the nature of each bead. These decoder probes light up through fluorescent die so that machines can “see” the sequence layout.
What Are Bead Arrays Used For?
Illumina uses bead arrays and oligonucleotide probes for genotyping DNA samples.
To learn more, check out our article that explains what genotyping is.
Companies like Ancestry.com and 23andMe use Illumina machines and genotyping to give consumers an analysis of their DNA.
Hello, I have been using direct to customer DNA testing for several years now and the information that C & G and A & T only combine surprises me and is new knowledge. I do phasing of kits because as you know the technology has yet to be developed to distinguish Paternal and Maternal contributions as far as the RAW data are concerned as supplied to the customer for up loading to other sites like Gedmatch etc. Missing parent phasing produces a 50% kit for the second parent, and an immediate application of this new knowledge is to run a program or function in Excel to pair the missing SNP per base when the missing parent kit has been produced. This will make up loading and comparisons much easier, as I use a look up table to do phasing based on what permutations are possible between a parent and child, I’ll now do some checking of my many kits to see how this works in practice, and how many errors in the measurements recorded are apparent.